Did you know that our genes can actually influence us to be more susceptible to coronaviruses? But how?
The common belief is that everyone has an equal chance of getting infected from coronaviruses upon exposure.
However, studies have suggested that genetic variations in immune function-related genes (E.g. human leukocyte antigens) are emerging as a critical determinant of COVID-19 infection[6]. Let’s find out more about our genes and understand how our immune system fights against coronaviruses.
READ ALSO: Project CoviDNA: Identify Your Coronavirus Infection Risk & Severity
What Is Coronavirus?
Coronaviruses are named for the crown-like spikes on their surfaces; they were first identified in the mid-1960s[3].
Type of coronaviruses
3 well-known coronaviruses that can infect people are[3]:
- MERS-CoV:
the beta coronavirus that causes Middle East Respiratory Syndrome or MERS - SARS-CoV:
the beta coronavirus that causes Severe Acute Respiratory Syndrome or SARS - SARS-CoV-2:
the novel coronavirus that caused the coronavirus disease 2019 or COVID-19
Gene Variant In Interleukins
What are interleukins?
Interleukins are a group of proteins that are made primarily in immune system cells. They are involved in cell-to-cell communication and have a wide variety of functions within the immune system. In response to stress-generating internal processes (e.g. cancer or microbial/virus infection), host cells secrete cytokines with a highly important role in cell metabolism reprogramming as a defensive response.
Key cytokines include those involved in adaptive immunity (e.g. IL-2 and IL-4), proinflammatory cytokines, interleukins (ILs) and anti-inflammatory.
Our immune system is made up of two armies of cells: innate immunity and adaptive/acquired immunity. Adaptive immunity is the body’s second line of defence against the pathogens[2]. It is activated by the exposure to pathogens and uses an immunological memory to learn about the threat and enhance the immune response accordingly[2].
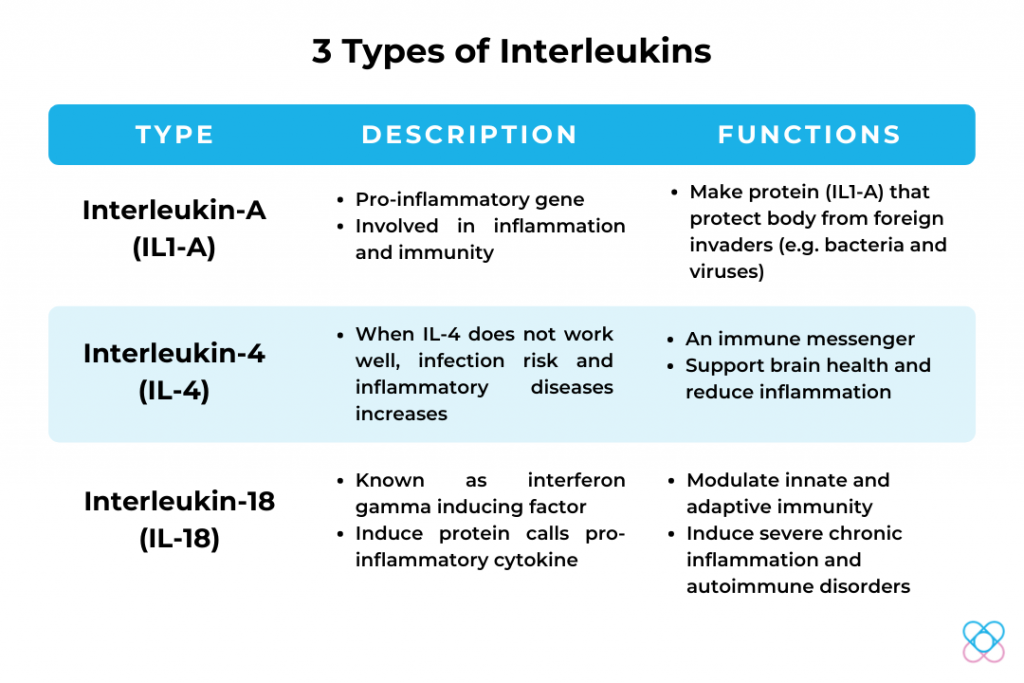
IL1-A
Individuals with genotype T of IL1-A have shown an association with SARS-CoV virus load[1,5]. Gene variant individuals of IL1-A show a higher risk of SARS-CoV virus load.
Virus load: The amount of virus in an infected person’s blood.
A study was conducted in a group of SARS-patients; the results showed that patients with high SARS-CoV virus load were associated with complicated clinical issues, such as acute respiratory distress syndrome (ARDS), which is known as respiratory failure[1,4].
IL-4
Individuals with genotype T of IL-4 have shown an association with respiratory infection[5]. A study reported that mild symptoms such as fever, followed by the development of respiratory symptoms, may progress to acute respiratory distress syndrome (ARDS) in some patients[1].
In addition, research was conducted in a group of SARS-patients with high initial or peak nasopharyngeal virus load that is associated with a high mortality rate[4]. It was also observed that patients with a high SARS-CoV virus load were associated with an increased risk for admission to an intensive care unit (ICU)[4].
IL-18
Interleukin-18 is an essential pro-inflammatory cytokine involved in autoimmune disorders and chronic inflammation, in responding to both innate and adaptive immunity[11].
Individuals with genotype T of IL-18 have shown an association of increased risk of virus shedding[4,5]. The research was done on a group of SARS-patients with the highest level of virus shedding showed a risk of early death throughout the infected period[4].
Virus shedding: The expulsion and release of virus (successfully virus replication in the body) from an infected individual. Throughout the process, individuals may or may not experience any viral symptoms (e.g. fever). It is normally transmitted via talking, eating, exhaling and performing normal daily activities (e.g. shopping, working).
Gene Variant In Human Leukocyte Antigens
What Are Human Leukocyte Antigens?
Human leukocyte antigens or HLAs, the human version of the major histocompatibility complex (MHC), genes that occur in many species[9]. Genes in this complex are categorised into 3 basic groups – MHC class I, II and III[9].
Basically, the HLAs complex act like a human body’s immigration officers. It helps the immune system to differentiate the proteins belonging either to the body itself or foreign invaders (e.g. bacteria, virus)[9,12]. Besides, it also helps the body to determine the outcomes of many infectious diseases such as HIV and SARS[12].
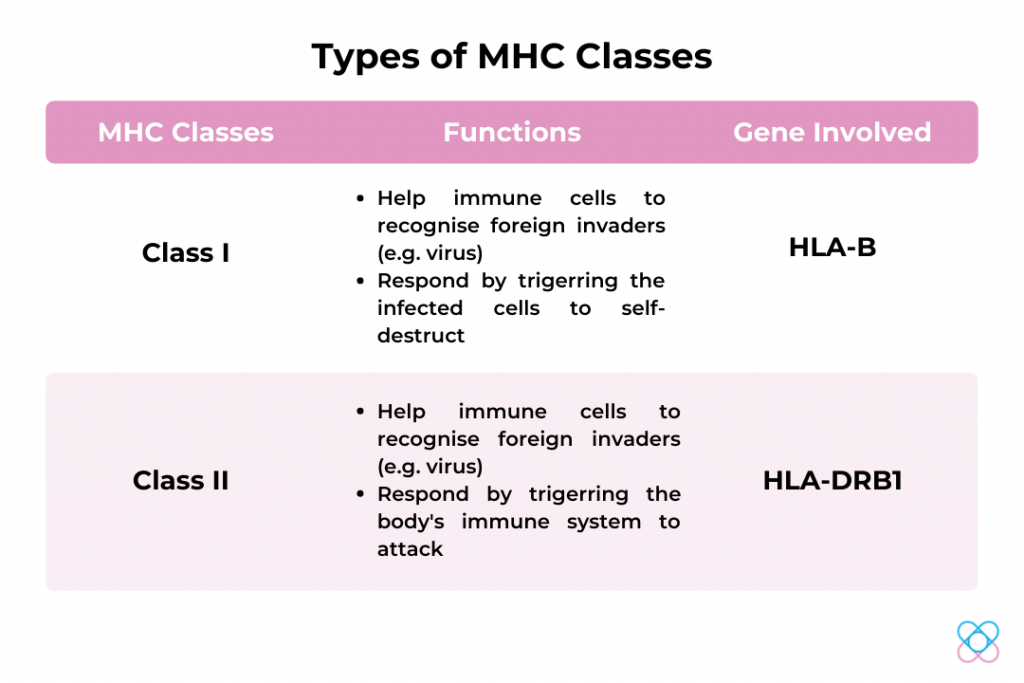

HLA-B
Genetic variation in HLA-B genes may be associated with the occurrence of SARS-CoV-2 infection [10,12]. Individuals with gene variants in the HLA-B gene might have a higher chance of contracting severe SARS-CoV- 2 infection.
In a study, a group of 33 patients with severe SARS-CoV- 2 infections were admitted into intensive care units (ICU) and 7 were deceased during the period[10].
HLA-DRB1
A study was done by Magaret et. al (2004), the HLA-DRB1 gene provides protection against infection with SARS[10]. Therefore, the HLA-DRB1 gene may have shown the possibility to be resistant to SARS-CoV infection.
Gene Variant In CD14
What is CD14?
CD14 (cluster of differentiation 14) is a surface antigen (receptor) which encodes a human protein that is preferentially expressed on white blood cells (e.g. monocytes)[7,13]. This protein plays a crucial role in immune recognition and reactivates microbial cell wall components from bacteria[7]. It also cooperates with other proteins to mediate the innate immune response to bacteria and viruses[13].
Innate immunity is one of the two armies of cells that make up our immune system. It is the first-line defence of the body virus[2]. They quickly respond to foreign cells to fight infection, battle a virus or defend the body against bacteria. For instance, physical barriers (e.g. hairs, skins), chemical barriers (e.g. mucus membrane, stomach acid) and cellular defence [2].
CD14
CD14 belongs to the innate immunity system; it is the key molecule to activate the innate immune cells. Individuals with genotype C of CD14 have shown an association of risk and severity with SARS-CoV disease[13].
A study reported that patients with high severity of SARS-CoV disease are either admitted to the intensive care unit (ICU) or are early deceased[13]. Besides, they also have weaker immune systems due to weak antiviral activity affected by the virus[13]. Hence, they may have higher chances to contract a more severe-form of SARS-CoV virus infection.
Furthermore, CD14 has been identified as a target candidate in the treatment of SARS-CoV-2-infected patients to potentially lessen or inhibit a severe inflammatory response[8]. However, more research still needs to be done in order to ensure the potency of CD14 as a target candidate for the treatment of SARS-CoV-2.
Is It The End?
According to the Centres for Disease Control and Prevention (CDC), there is currently no specific treatment for coronaviruses. To prevent contracting coronaviruses, the current preventive methods are to wash your hands with soap, avoid touching your eyes or nose with unclean hands, as well as to clean and disinfect frequently touched surfaces and objects (e.g. table and doorknob).
At this point, our knowledge of the relationship of coronaviruses and immune systems is limited. Scientists have been thoroughly studying the relationship between SARS, MERS and COVID-19.
Although vaccination has been undergoing clinical trials (which will be internationally available next year), the genome sequence of coronaviruses is yet to be completely discovered and well-studied. Hence, more updates on research and data will be available in the future. By then, we hope to have a clearer understanding of the relationship between these three viruses.
For now, let’s do our part by washing our hands, wear our masks, social distance, and practice all the necessary safety precautions.

Handwash with Soap 
Social Distance 
Wear Mask
Find out your genetic predisposition in infection risk and severity when exposed to coronaviruses with Project CoviDNA today!
References
1. Badraoui, R., Alrashedi, M. M., El-May, M. V., & Bardakci, F. (2020). Acute respiratory distress syndrome: a life threatening associated complication of SARS-CoV-2 infection inducing COVID-19. Journal of Biomolecular Structure and Dynamics, 1–10. https://doi.org/10.1080/07391102.2020.1803139
2. Cancer Treatment Centers of America. (2017). What’s the Difference between B-Cells and T-Cells. Retrieved 2 December 2020, from https://www.cancercenter.com/community/blog/2017/05/whats-the-difference-b-cells-and-t-cells
3. Centers for Disease Control and Prevention. (2020). Coronavirus | Human Coronavirus Types | CDC. Retrieved 7 December 2020, from https://www.cdc.gov/coronavirus/types.html
4. Chen, W.-J., Yang, J.-Y., Lin, J.-H., Fann, C. S. J., Osyetrov, V., King, C.-C., Chen, Y.-M. A., Chang, H.-L., Kuo, H.-W., Liao, F., & Ho, M.-S. (2006). Nasopharyngeal Shedding of Severe Acute Respiratory Syndrome–Associated Coronavirus Is Associated with Genetic Polymorphisms. Clinical Infectious Diseases, 42(11), 1561–1569. https://doi.org/10.1086/503843
5. Di Maria, E., Latini, A., Borgiani, P., & Novelli, G. (2020). Genetic variants of the human host influencing the coronavirus-associated phenotypes (SARS, MERS and COVID-19): rapid systematic review and field synopsis. Human Genomics, 14(1). doi: 10.1186/s40246-020-00280-6
6. Debnath, M., Banerjee, M., & Berk, M. (2020). Genetic gateways to COVID‐19 infection: Implications for risk, severity, and outcomes. The FASEB Journal, 34(7), 8787-8795. doi: 10.1096/fj.202001115r
7. Fatemeh, Z., Fateme, Z., Leili, A., & Behzad, B. (2013). Induction of CD14 Expression and Differentiation to Monocytes or Mature Macrophages in Promyelocytic Cell Lines: New Approach. Advanced Pharmaceutical Bulletin, 3(2), 329-323. doi: 10.5681/apb.2013.053
8. National Center for Biotechnology Information. (2020). CD14 molecule [Homo sapiens (human)] – Gene. Retrieved 8 December 2020, from https://www.ncbi.nlm.nih.gov/gene/929
9. National Institute of Health. (2020). HLA-B gene: MedlinePlus Genetics. Retrieved 9 December 2020, from https://medlineplus.gov/genetics/gene/hla-b/
10. Ng, M., Lau, K., Li, L., Cheng, S., Chan, W., & Hui, P. et al. (2004). Association of Human‐Leukocyte‐Antigen Class I (B*0703) and Class II (DRB1*0301) Genotypes with Susceptibility and Resistance to the Development of Severe Acute Respiratory Syndrome. The Journal Of Infectious Diseases, 190(3), 515-518. doi: 10.1086/421523
11. Sánchez, E., Palomino-Morales, R., Ortego-Centeno, N., Jiménez-Alonso, J., González-Gay, M., & López-Nevot, M. et al. (2009). Identification of a new putative functional IL18 gene variant through an association study in systemic lupus erythematosus. Human Molecular Genetics, 18(19), 3739-3748. doi: 10.1093/hmg/ddp301
12. Wang, W., Zhang, W., Zhang, J., He, J., & Zhu, F. (2020). Distribution of HLA allele frequencies in 82 Chinese individuals with coronavirus disease‐2019 (COVID‐19). HLA, 96(2), 194-196. doi: 10.1111/tan.13941
13. Yuan, F. F., Boehm, I., Chan, P. K. S., Marks, K., Tang, J. W., Hui, D. S. C., Sung, J. J. Y., Dyer, W. B., Geczy, A. F., & Sullivan, J. S. (2007). High Prevalence of the CD14-159CC Genotype in Patients Infected with Severe Acute Respiratory Syndrome-Associated Coronavirus. Clinical and Vaccine Immunology, 14(12), 1644–1645. https://doi.org/10.1128/cvi.00100-07

